Effect of frame numbers and analysis algorithms on superresolution images
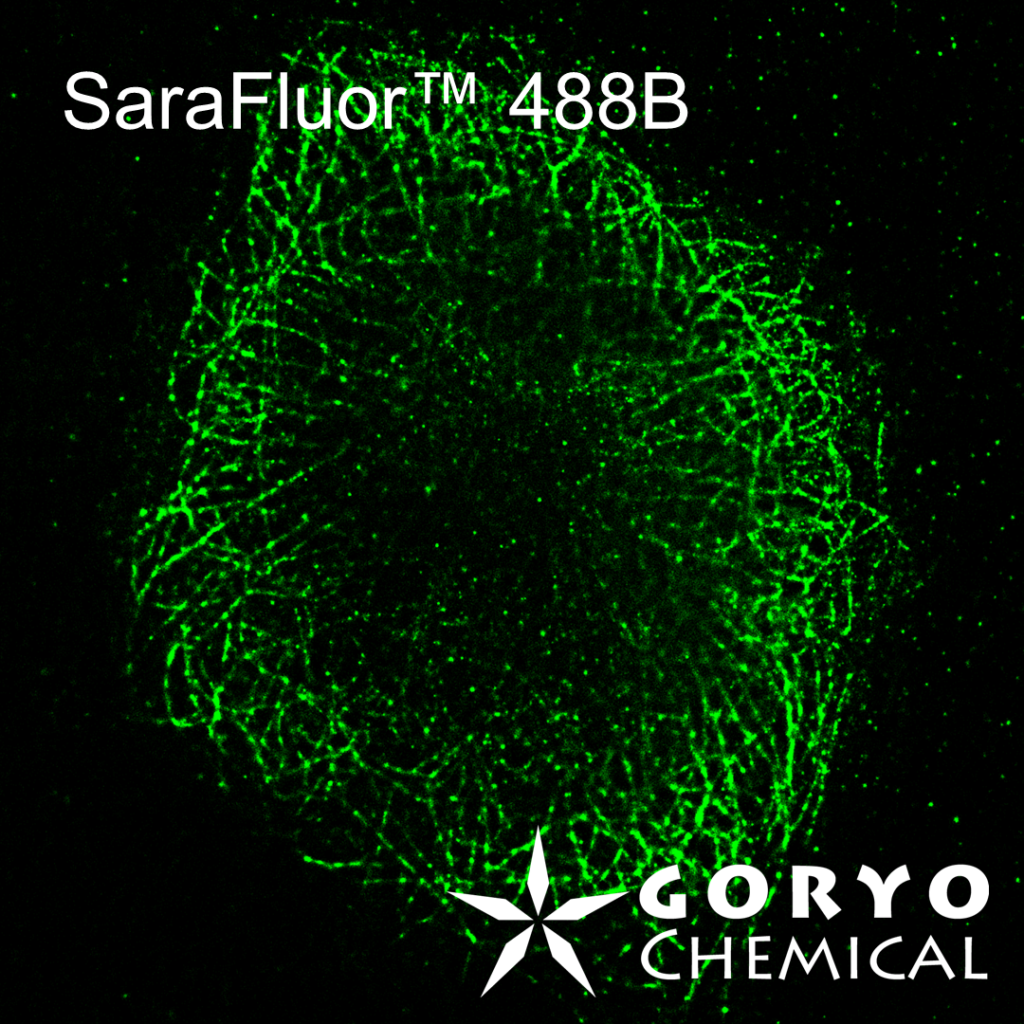
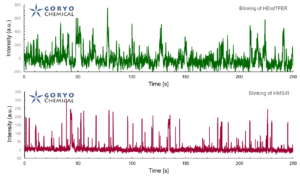
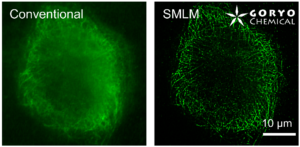
SaraFluor B series is a line of fluorophores which show spontaneous blinking in physiological pH. Without a special microscope system for PALM/STORM, a microscope which can image single fluorophore can be used to get multiple images of SaraFluor B and these images can be converted to superresolution image by using ImageJ plugins.
Besides a commonly used ImageJ plugin, ThunderSTORM, TRM mode of NanoJ SRRF , a plugin for supreresolution image analysis, can be also used to analyze SaraFluor B images to obtain superresolution images. Here, we qualitatively compare the effect of image numbers using these two plugins.
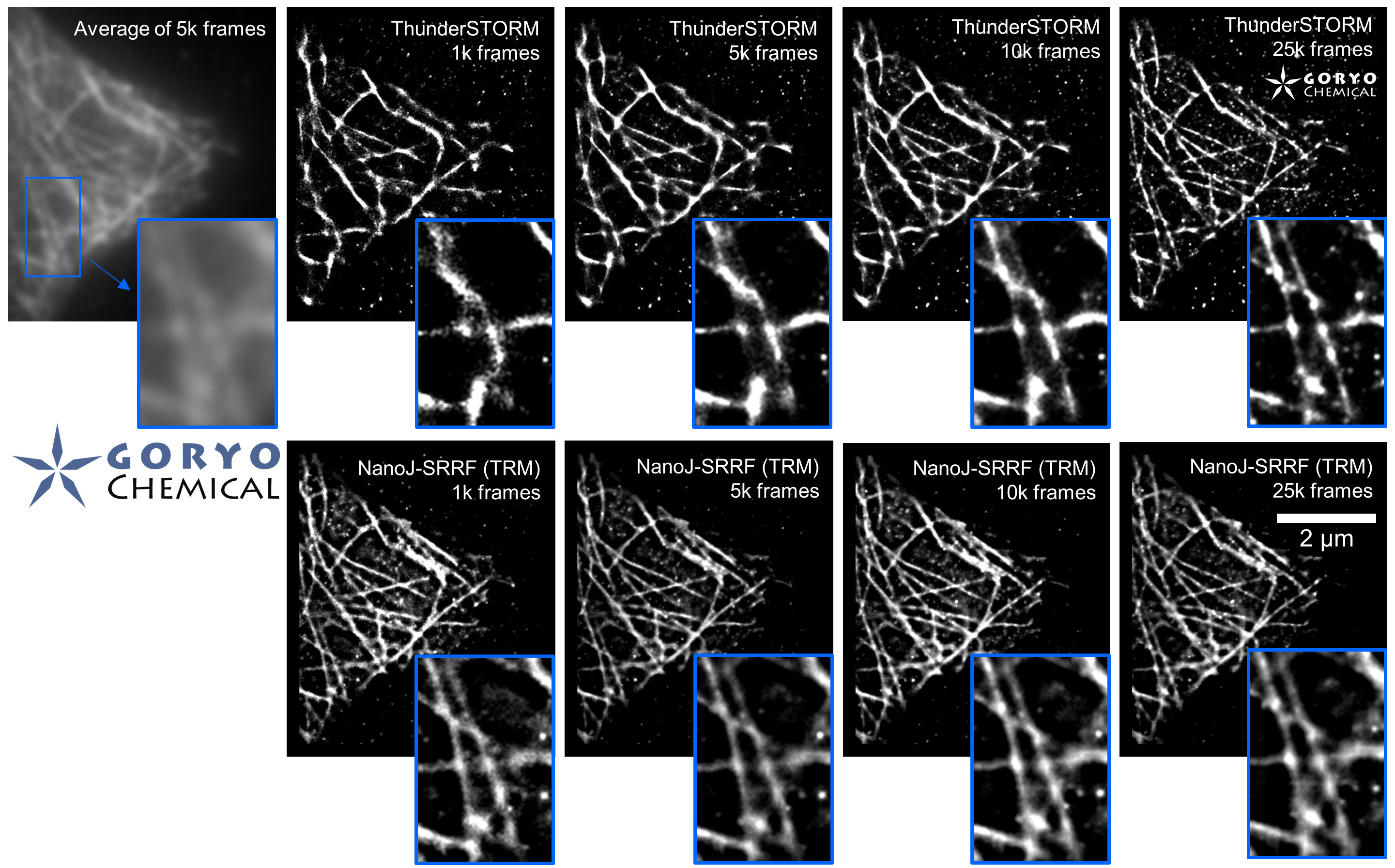
Fixed HeLa cells were reacted with anti-α tubulin antibody (DM1A, 1/4000) as a primary antibody, and then reacted with SaraFluor™ 650B goat anti-mouse IgG (A202-01, 30 μg/mL) as a secondary antibody. The cells were observed with NIKON N-STORM system by TIRF illumination using 647-nm laser, 100x Apo TIRF (NA 1.49) lens . Exposure was 30 msec/frame. Images were analyzed by the each softwares.
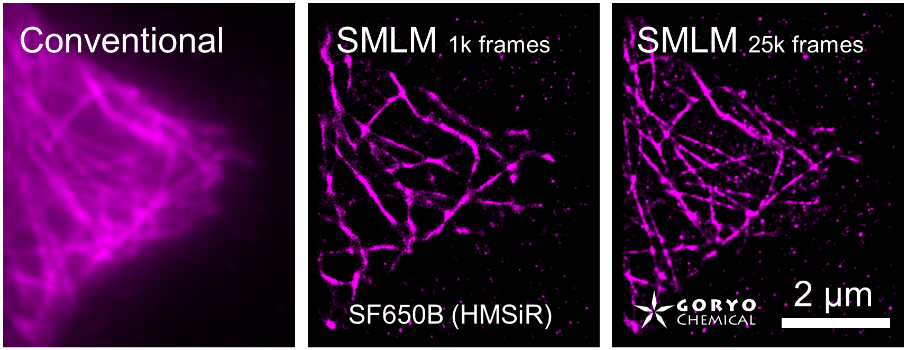
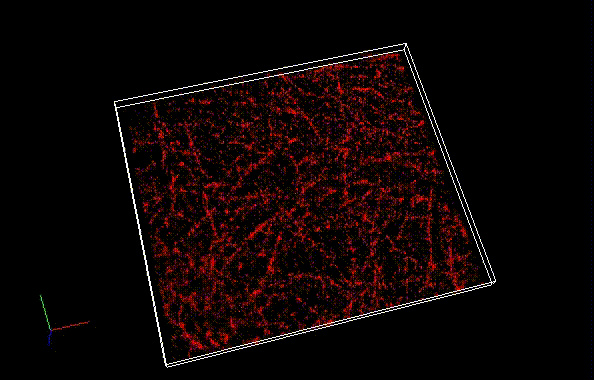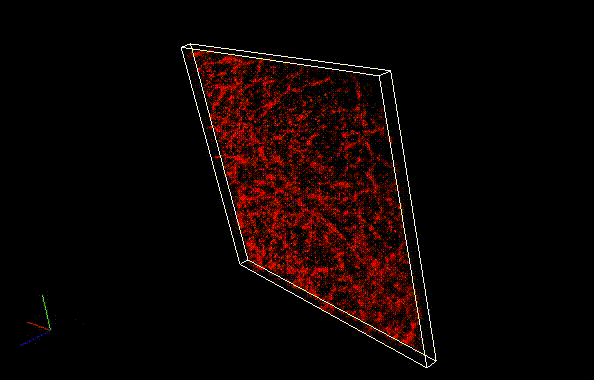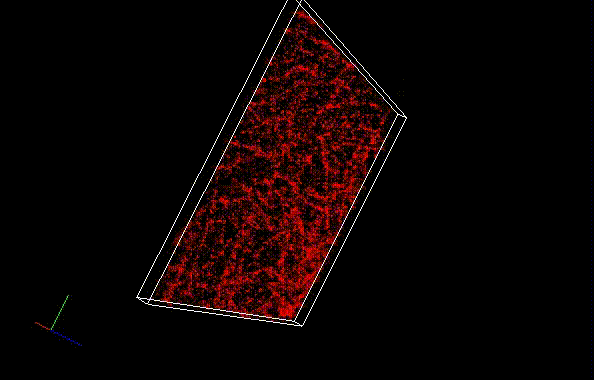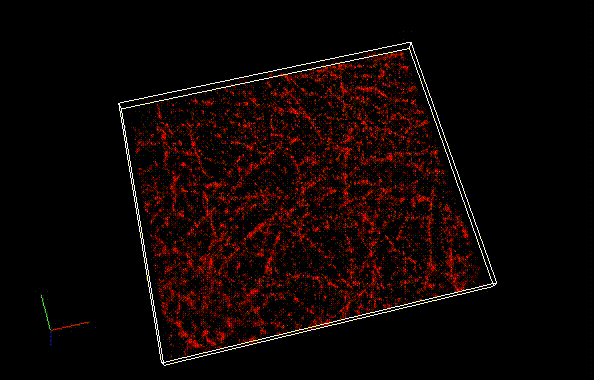
By using ThunderSTORM, 1,000 images were not enough to image some of the microtubule structures, whereas 10,000 images seems to be enough to image most of the microtubules. NanoJ SRRF (TRM) shows more information for fewer image numbers. ThunderSTORM provided higher contrast, whereas NanoJ SRRF did not give higher contrast and higher resolution even when using >10,000 images.
References
M. Ovesný, P. Křížek, J. Borkovec, Z. Švindrych, G. M. Hagen. (2014) Bioinformatics 30:2389-2390 DOI: 10.1093/bioinformatics/btu202 (ThunderSTORM)
Nils Gustafsson, Siân Culley, George Ashdown, Dylan M. Owen, Pedro Matos Pereira, & Ricardo Henriques (2016) Nature Communications 7: 12471 DOI: 10.1038/ncomms12471 (NanoJ SRRF)
Acknowlegements
We appreciate Dr. Kentaro Kobayashi and Dr. Motosuke Tsutsumi (NIKON Imaging Center, Hokkaido Univ.) for advices on imaging and analysis.









 Contact Us
Contact Us
